ZJU scientists identify sDscams as a candidate to mediate cell recognition in Chelicerata
The research team headed by Prof. JIN Yongfeng from the Zhejiang University College of Life Sciences published an article entitled “Chelicerata sDscam isoforms combine homophilic specificities to define unique cell recognition” in the September 22 issue of PNAS.
Neuronal self-avoidance, as a fundamental feature in vertebrates and invertebrates, plays a crucial role in neural circuit assembly.In Drosophila, self-avoidance is mediated by the Down syndrome cell adhesion molecule (Dscam1) gene that encodes tens of thousands of proteins through alternative splicing. In sharp contrast to insect Dscam1, vertebrate Dscam genes do not produce extensive isoform diversity. In vertebrates, a different set of cell surface adhesion receptors, the cPcdhs, perform an analogous function.
In 2016, JIN Yongfeng et al. discovered a “hybrid” gene family in the subphylum Chelicerata that is particularly relevant to the remarkable functional convergence of Drosophila Dscam1 and vertebrate cPcdhs. This gene family is composed of Dscam-related genes with tandemly arrayed 5′ cassettes, which encode ∼50 to 100 isoforms each with alternative promoters for the number of isoforms varying across Chelicerata species. Although these Chelicerata Dscams are evolutionarily related to Drosophila Dscam1, they only have 6 extracellular domains, making them much shorter compared to the 16 domains of Drosophila Dscam1. This type of Dscam is therefore referred to as shortened Dscam (sDscam). However, little is known about the Dscam1-related family of ∼100 sDscam proteins in Chelicerata.
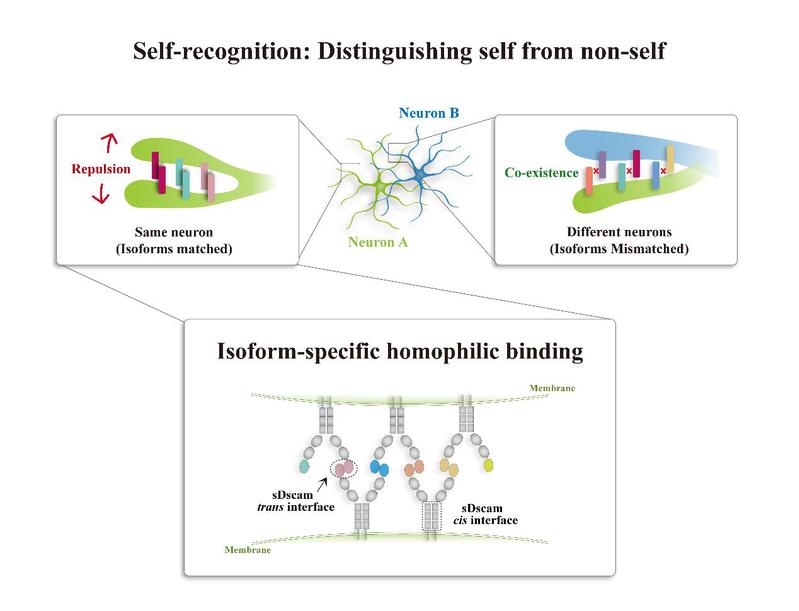
In the current research, JIN Yongfeng et al. find that Chelicerata sDscamα and some sDscamβ protein trans interactions are strictly homophilic, and that the trans interaction is meditated via the first Ig domain through an antiparallel interface. Additionally, different sDscam isoforms interact promiscuously in cis via membrane proximate fibronectin-type III domains. They also discover that cell–cell interactions depend on the combined identity of all sDscam isoforms expressed. A single mismatched sDscam isoform may well interfere with the interactions of cells that otherwise express an identical set of isoforms. Thus, these data underpin a model by which sDscam association in cis and trans generates a vast repertoire of combinatorial homophilic recognition specificities. Researchers thus propose that in Chelicerata, sDscam combinatorial specificity can provide each neuron with a unique identity for self–nonself discrimination. Surprisingly, while sDscams are related to Drosophila Dscam1, their results are consistent with the findings reported for the structurally unrelated vertebrate cPcdh.
This study serves as a remarkable example of convergent evolution for the process of neuronal self-avoidance and provides insights into the basic principles and evolution of metazoan self-avoidance and self–nonself discrimination.