ZJU scientists discover physiological function of CNOT4 subunit in XY chromosome crossover during male germ cell meiosis
On March 16, the lab led by FAN Hengyu from the Zhejiang University Life Sciences Institute published an open-access article entitled “The CNOT4 Subunit of the CCR4-NOT Complex is Involved in mRNA Degradation, Efficient DNA Damage Repair, and XY Chromosome Crossover during Male Germ Cell Meiosis” in the journal Advanced Science. It is the first time that researchers have reported the physiological function of the CNOT4 subunit of the CCR4-NOT complex during the process of embryo development and meiosis progression during spermatogenesis and proposed that CNOT7 has CNOT6/6L-independent functions in vivo.
Previous studies in gene knockout mice indicated that the two subunits of the CCR4-NOT complex—CNOT7 and CNTO6L—played crucial role in the development of male and female germ cells, but there is a paucity of research into the physiological function of other subunits which play synergistic biochemical functions with them.
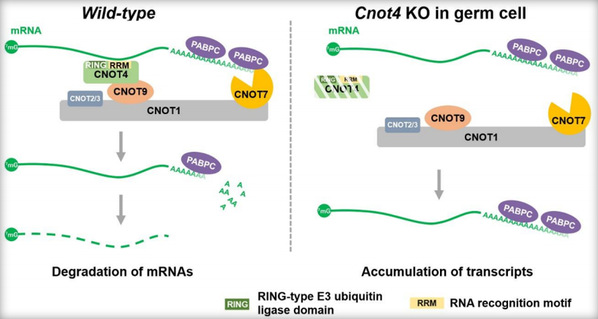
In this study, researchers first knocked out both Cnot6 and Cnot6l in mice. Surprisingly, Cnot6/6l-/- mice were viable and healthy, indicating that Cnot6/6l is not essential for survival but responsible for regulating oogenesis. Meanwhile, experiments revealed that Cnot4 knockout was lethal to embryos, suggesting the more universal physiological function of this subunit. Conditional knockout of Cnot4 in male germ cells led to defective spermatogenesis and male infertility. These findings provided hereditary evidence under physiological conditions, indicating that CNOT7 can play a vital role independently of CNOT6/6L in male germ cells and that CNOT4 functions as a previously unrecognized mRNA adaptor of CCR4-NOT by targeting mRNAs to CNOT7 for deadenylation of poly(A) tails, thereby mediating the degradation of a subset of transcripts from the zygotene to pachytene stage. These findings disproved the idea that CNOT6/6L and CNOT7/8 should play synergistic roles in deadenylation of poly(A) tails.

Researchers probed into the function of the CNOT4 subunit of the CCR4-NOT complex during meiosis in male germ cells and confirmed that at the physiological level CNOT4 is an RNA-binding protein that acts as a substrate recognition subunit. During meiosis, CNOT4 can regulate the normal degradation of mRNA during the zygotene-to-pachytene transition through an RNA recognition motif (RRM). This biochemical process fulfills several functions as follows. First, it can promote the appropriate expression of genes in spermatogenesis. Second, it can ensure the normal DNA double-strand break repair during meiosis. Third, it can contribute to the efficient crossover between X and Y chromosomes.
In this study, researchers pioneered in establishing links between the regulation of mRNA stability and other important events in reproductive biology, such as DNA break repair during meiosis and efficient crossover between X and Y chromosomes. This study will open the door to new research molecular and developmental biology.