Scientists probe into spinach domestication and genetic basis of agronomic traits
On December 13, Prof. JIAO Chen from the College of Agriculture and Biotechnology, Zhejiang University, and Prof. FEI Zhangjun from the Boyce Thompson Institute, Cornell University, co-published a research article entitled “Genomic analyses provide insights into spinach domestication and the genetic basis of agronomic traits” in the journal Nature Communications.
Spinach is a remarkably nutritious leafy vegetable belonging to the Chenopodiaceae family. However, there has been a paucity of research into its resistance to major diseases and its improvement, due largely to lack of data regarding its genome and genetic diversity. To this end, JIAO Chen and collaborators conducted relevant research into the draft genome of spinach. Their research findings were previously published in the journal Nature Communications in 2017. This genome is still in the draft phase owing to the limitations of technology and material.
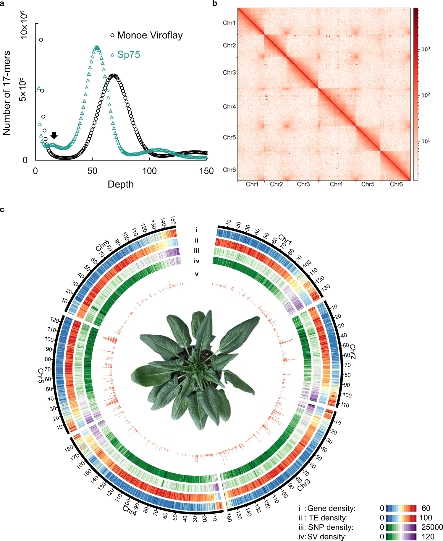
Genome of the spinach Monoe-Viroflay
In the present study, the researchers assembled a much-improved chromosome-scale spinach reference genome using PacBio long reads and the high-throughput chromosome conformation capture (Hi-C) technology. They reconstructed the ancestral karyotype of Chenopodiaceae, which indicated substantial genome rearrangements in spinach after its divergence from ancestral Chenopodiaceae, coinciding with high repeat content in the spinach genome.
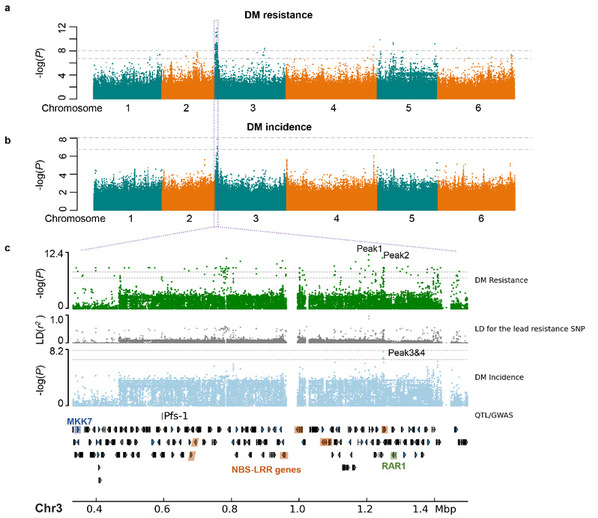
GWAS of spinach downy mildew (DM) resistance
Jiao and collaborators sequenced the genomes of 305 cultivated and wild spinach accessions collected worldwide and constructed a high-density genomic variation map which allowed them to dissect the genetic architecture of complex spinach traits using genome-wide association studies (GWAS). Meanwhile, their GWAS analyses on 20 agronomic traits identified a series of genomic regions associated with these traits and candidate genes, thereby facilitating the breeding of spinach cultivars featured by downy mildew resistance and reduced soluble oxalate content. In addition, by comparing the genetic diversity of wild and cultivated spinach, the researchers also identified the genomic domestication characteristics of spinach differentiated from other crops as well as some selected regions associated with spinach domestication traits (e.g. leaf phenotype, bolting and flowering), demonstrating the role of artificial selection in shaping spinach genomic and phenotypic evolution.
“Our study provides not only insights into spinach evolution and domestication but also valuable resources for facilitating spinach breeding,” said Jiao.