Optimization of metabolomic data processing using machine learning and parallel computing
Metabolomics involves the qualitative and quantitative analysis of the unique chemical fingerprints that specific physiological and pathological processes leave behind. Due to its uniqueness of closely linking to the phenotypes, metabolomics is applied to a variety of stages in drug discovery (such as drug target identification, lead compound discovery, drug metabolism analysis, drug response and resistance), and expected to significantly accelerate the development of new drug. However, the successful application of current metabolomic technique is severely limited by both signal processing and data analysis. To effectively remove the unwanted environmental, instrumental and biological signal flucturations, it is urgently needed to develop novel methods for identifying the optimal data processing strategy tailored for different metabolomic studies.

Recently, a research team led by Professor ZHU Feng at the College of Pharmaceutical Sciences of Zhejiang University conducted a collaborative study with Alibaba-Zhejiang University Joint Research Center of Future Digital Healthcare. Their research findings were published in the Nature Protocols to address those critical issues described above, and highlighted as a Featured Article on journal’s official website. In their pioneer study, NOREVA, a novel and out-of-the-box software tool capable of determining the optimal processing strategy for a specific metabolomic study, was developed.
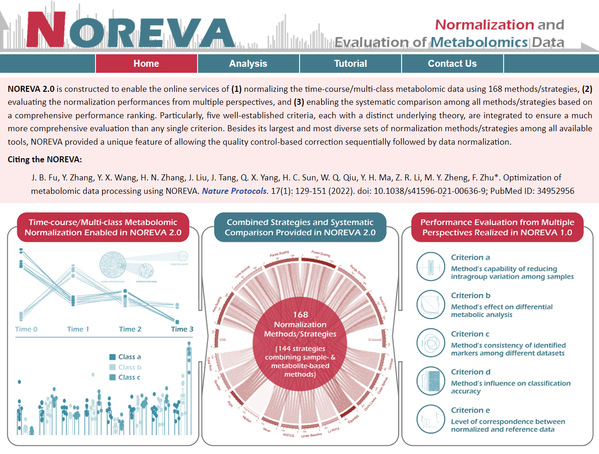
In NOREVA, peak table was first subjected to many processing workflows comprised of three to five defined calculations in combinatorially determined sequences. Then, the results of each workflow were evaluated against multiple objective performance criteria. Third, benchmarks were analyzed to highlight the uniqueness of this newly developed tool in evaluating and optimizing thousands of workflows in a single experiment, and allowing data processing for both time-course and multiclass metabolomics. These uniquenesses make NOREVA an indispensable tool in modern target identification, lead discovery, drug metabolism analysis, drug response, drug resistance, and so on.
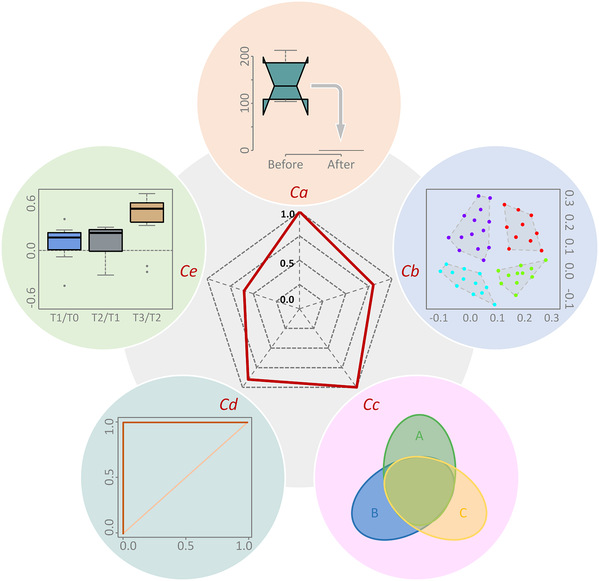
To address the bottleneck problem regarding computational resources during the large-scale scanning of massive signals and processing workflows, this study introduced the parallel computing, for the first time, to metabolomics. “Trials in our research showed that in contrast with serial computing, parallel computing could substantially improve operational efficiency more than 10-folds on personal computers,” said Prof. Zhu. “We are currently promoting the deployment and offering services on AliCloud.”
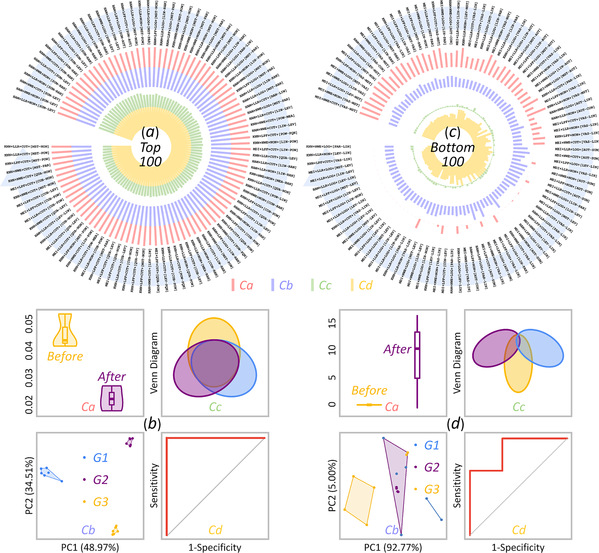
More information: Dr. FU Jianbo and Dr. ZHANG Ying are co-first authors of this study.