Viral relics in the genome help human embryos switch on their own genes, ZJU scientists find
A large fraction of the human genome is built from the remnants of ancient viral infections. Long dismissed as genetic clutter, these sequences are increasingly being implicated in early development. Now a team in China reports that a sub-family of these elements may help human embryos clear one of their earliest and most precarious milestones.
In a study published in Science on January 22, researchers led by LIANG Hongqing at Zhejiang University’s School of Basic Medical Sciences and Women’s Hospital of Zhejiang University School of Medicine, together with ZHANG Dan at Women’s Hospital, GAO Shaorong at Tongji University and LIU Wanlu at the ZJU–UoE Institute, describe how endogenous retroviruses (ERVs) can generate unusual chimeric RNAs that appear to bolster zygotic genome activation (ZGA), the moment when an embryo begins to transcribe its own genome rather than relying on maternal RNAs and proteins. The work, the authors say, could eventually benefit embryo assessment in assisted reproduction, although clinical translation remains a distant goal.
A clinical puzzle meets genomic “dark matter”
In in vitro fertilization (IVF) clinics, embryo arrest before implantation is a stubborn and often unexplained problem. “In clinical practice we see many IVF failures,” ZHANG Dan says. “We conduct comprehensive checks of patients’ chromosomes and protein-coding genes, yet still can’t find the cause.” For clinicians and patients, these cases can feel like biological dead ends.

The project took shape when ZHANG Dan exchanged ideas with LIANG Hongqing, whose lab studies transposable elements — repetitive sequences that populate the genome but rarely encode proteins. If conventional genetic screening comes up empty, the researchers reasoned, the explanation might lie outside the coding regions.
Transposable elements make up a substantial chunk of the human genome, and ERVs are one of the most prominent class. Over the course of evolution, these viral sequences have been co-opted by their hosts and can influence gene regulation. Yet in early human embryos, where ERV-derived RNAs surge in a stage-specific manner, their functions have been difficult to pin down.
A signal at the 8-cell stage
ZGA is a dramatic handover. Before it, the embryo is largely governed by maternal RNAs and proteins deposited in the egg; after it, the embryo must begin running its own genetic program. Failures in this transition are thought to contribute to early developmental arrest.
The team reports that in 8-cell embryos that stopped developing after IVF — a stage widely used in assisted reproduction as a checkpoint for embryo evaluation, expression of an ERV subfamily called MLT2A1 was markedly reduced. In the authors’ experiments, lower MLT2A1 levels were closely bound up with reduced expression of genes associated with ZGA, hinting that the element might be doing more than simply “switching on” alongside development.
To probe what MLT2A1 produces, the researchers used nanopore long-read sequencing in human embryos and stem cells. What emerged was not a single transcript, but hundreds of distinct RNA variants: MLT2A1 sequences fused to different downstream genomic regions, forming a diverse set of MLT2A1 chimeric RNAs. Rather than acting as a uniform transcript, MLT2A1 appears to generate a repertoire and that, variety, the team argues, may be the point.

“During ZGA, MLT2A1 is not merely transcribed as a genetic sequence,” ZHANG Dan says. Instead, it “fuses with various downstream genomic sequences,” producing many structurally diverse chimeras.
If chimeric RNAs are functional, they should interact with the genome in a targeted way. Using ChIRP (Chromatin Isolation by RNA Purification), the researchers mapped where MLT2A1 chimeric RNAs associate across DNA. They report binding at key regulatory regions, including gene promoters, as well as other subfamilies of retro-transposable elements.
Crucially, different chimeric sequences appeared to align with activation of different ZGA genes. In the authors’ framing, the embryo does not rely on a single molecular switch but on a distributed set of RNA “adapters” that can plug into multiple sites, collectively supporting the global ramp-up in transcription that defines ZGA.
MLT2A1 RNA as a “coordinator”
How might RNA molecules influence genome-wide transcription in a system long thought to be protein-led? The team reports that MLT2A1 chimeras are enriched in the nucleus and form distinct condensate-like foci. Mechanistic experiments suggest these RNAs interact with the nuclear protein HNRNPU, promoting condensation and, in doing so, helping recruit RNA polymerase II, the enzyme that transcribes DNA into RNA.

The researchers tested the idea by perturbing the system. When they knocked down MLT2A1 or HNRNPU, newly synthesized RNAs from ZGA genes fell sharply and embryos showed developmental arrest. LIANG Hongqing describes MLT2A1 RNA as a “coordinator” that does two jobs at once: pairing with specific DNA sites while drawing transcriptional machinery into the neighborhood to boost gene activation.
“This MLT2A1–HNRNPU–RNA Pol II synergy provides a strong molecular basis for the global transcriptional activation event of zygotic genome activation,” ZHANG Dan says.
One feature of ERVs is their copy number: the human genome contains many near-repeats scattered across chromosomes. LIANG Hongqing notes that there are 3,838 copies of MLT2A1 distributed at different genomic sites, and that their chromatin accessibility and transcriptional activity peak during ZGA.
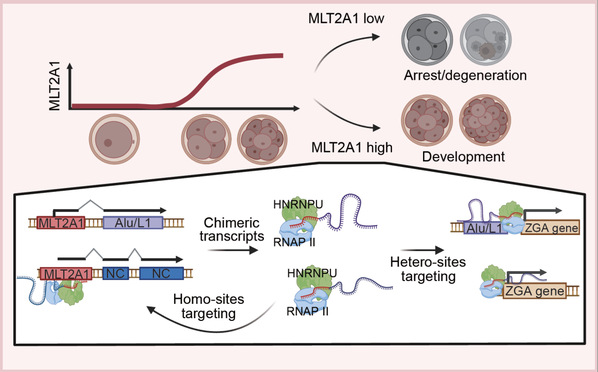
MLT2A1 reinforces human ZGA by synthesizing heterologous chimeric RNAs. ZGA-specific MLT2A1 generated chimeric RNAs by fusing with diverse downstream sequences.
The team reports a network-like behavior: disrupting one MLT2A1 copy not only reduces that copy’s transcript but also affects expression across other MLT2A1 elements, lowering the subfamily’s overall output. Conversely, overexpressing a single intact MLT2A1 chimeric RNA can boost expression of the subfamily more broadly. The authors interpret this as self-amplification and cooperation within the subfamily, potentially enabling a rapid, synchronized push during a developmental window where timing is everything.
What it could mean for IVF
The study adds to a growing picture in which transposable-element-derived RNAs are not passive byproducts but active participants in early development. It also offers a plausible molecular handle on a clinically important failure point: if ZGA does not initiate properly, embryos may stall before implantation.
The authors suggest that MLT2A1 chimeric RNAs could, in time, serve as a biomarker for embryo developmental potential in IVF. ZHANG Dan notes that measuring MLT2A1 expression might help clinicians identify embryos more likely to thrive after transfer, potentially improving pregnancy success rates. A deeper grasp of MLT2A1’s regulatory mechanism, the researchers add, could also open up new avenues to infertility linked to ZGA failure.

For now, the work offers something rarer than a quick fix: a new way to think about the earliest steps of human development and a reminder that, in biology, some of the most consequential signals can come from the genome’s most underestimated corners.
Translator: FANG Fumin
Editor: HAN Xiao