ZJU scientists develop multi-fidelity modeling for pKa prediction
Acid-base dissociation constant (pKa), a key physicochemical parameter describing the ionization ability of molecules, plays a vital role in chemical, environmental, agricultural, and medical sciences. In drug discovery, pKa determines the ionization states of drug-like molecules in the physiological environment, thus affecting a wide range of biopharmaceutical properties.
Graph Neural Networks (GNN) are a set of deep learning (DL) algorithms that can learn representation automatically from graph-structured data for node-, subgraph- or graph-level tasks, and have achieved tremendous success in molecular property predictions. However, due to the complexity of pKa, the applications of GNN in pKa predictions are relatively rare, even though they show immense promise.
The research team led by Prof. HOU Tingjun and Hsieh Chang-yu at the Zhejiang University College of Pharmaceutical Sciences developed MF-SuP-pKa (multi-fidelity modeling with subgraph pooling for pKa prediction), a novel pKa prediction model that utilizes subgraph pooling, multi-fidelity learning and data augmentation. Their research findings are published in an article titled “MF-SuP-pKa: Multi-fidelity modeling with subgraph pooling mechanism for pKa prediction” in the journal Acta Pharmaceutica Sinica B.
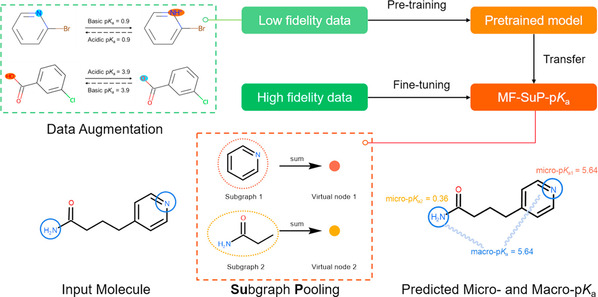
Graphical abstract
In this model, the researchers drew up a knowledge-aware subgraph pooling strategy to capture the local and global environments around the ionization sites for micro-pKa prediction. To address the scarcity of accurate pKa data, low-fidelity data (computational pKa) was employed to fit high-fidelity data (experimental pKa) through transfer learning. The final MF-SuP-pKa model was constructed by pre-training on the augmented ChEMBL data set and fine-tuning on the DataWarrior data set. Extensive evaluation of the DataWarrior data set and three benchmark data sets reveals that MF-SuP-pKa outperformed other state-of-the-art pKa prediction models while calling for much less high-fidelity training data. Compared with Attentive FP, MF-SuP-pKa improved its accuracy rate by 23.83% and 20.12% in terms of mean absolute error (MAE) on the acidic and basic sets, respectively.
Overall, this model is expected to serve as a powerful tool for the prediction of such biopharmaceutical properties as absorption, distribution, metabolism, excretion, and toxicity.





