AI-built “exoskeletons” offer a new way to steer GPCR signaling
G protein–coupled receptors (GPCRs) are among the most important “signal receivers” in the human body, translating the external stimuli into internal signals that govern sensation, mood, cardiovascular function and metabolism. These seven–transmembrane-helix proteins are also pharma favorites. More than a third of marketed drugs act by nudging GPCR activity. But many therapies still aim at a familiar target region: the orthosteric pocket where natural ligands bind, and that approach can struggle when disease arises from structural defects rather than a missing “on” signal.
Now, researchers at Zhejiang University report an unconventional approach: rather than pushing the receptor’s canonical “start button”, they wrap it in an engineered, membrane-embedded scaffold that constrains its motion and reshapes its output. ZHANG Yan’s team at the School of Medicine and the Liangzhu Laboratory worked with ZHANG Min’s team at the College of Computer Science and Technology, proposed de novo–designed “GPCR Exoframe Modulators” (GEMs) that function like a modulatory exoskeleton.
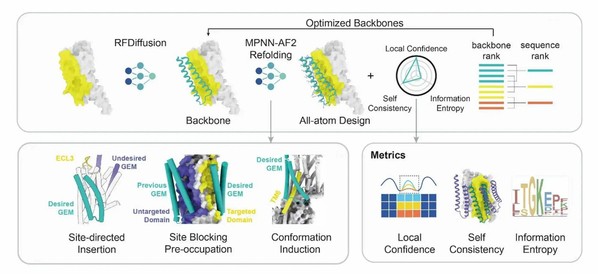
De novo Design of GEM.
During signaling, GPCRs undergo a distinctive seesaw-like conformational shift, converting external stimuli into intracellular responses. Traditional drug discovery typically aims to trigger or block this process by occupying the orthosteric pocket. “You can think of it as the receptor’s start button,” says ZHANG Yan. But that framing exposes a limitation. In clinical practices, hundreds of mutation-linked disorders can distort GPCR structures, leaving receptors misaligned, jammed or overly active. Drugs tuned to the orthosteric site often cannot restore proper mechanics, contributing to modest efficacy in some settings.
A further puzzle motivated the new work. Humans have more than 800 GPCRs, yet only a handful of naturally occurring transmembrane proteins are known to directly modulate the GPCR function. That mismatch hints at a large regulatory landscape on GPCR transmembrane surfaces that biology uses sparingly or that researchers have simply not learned to exploit.
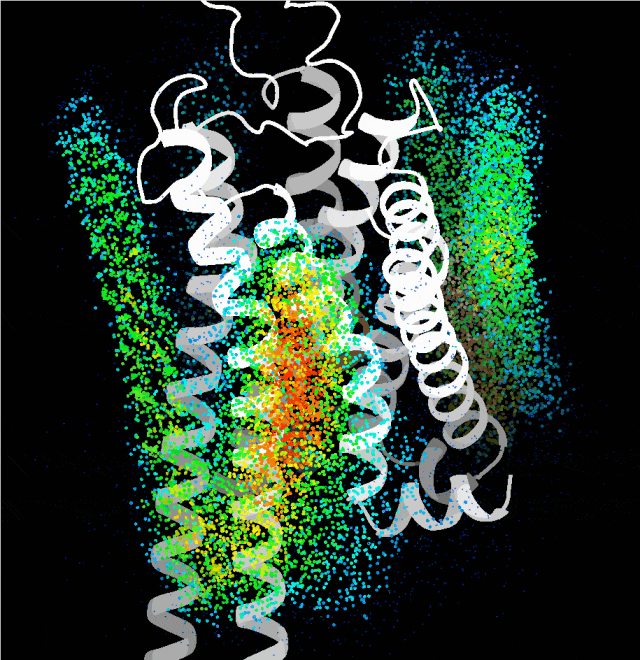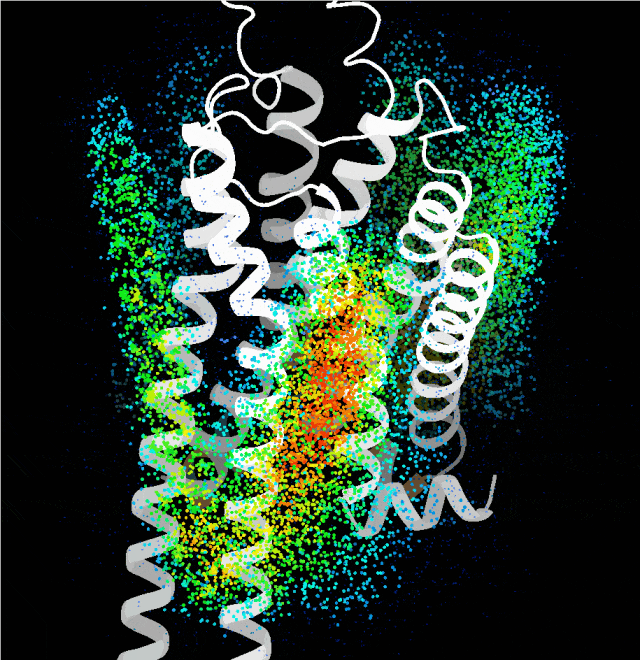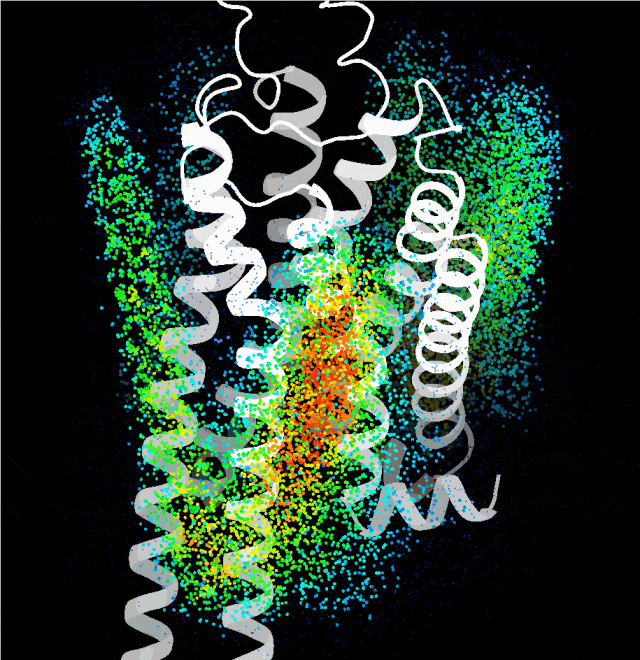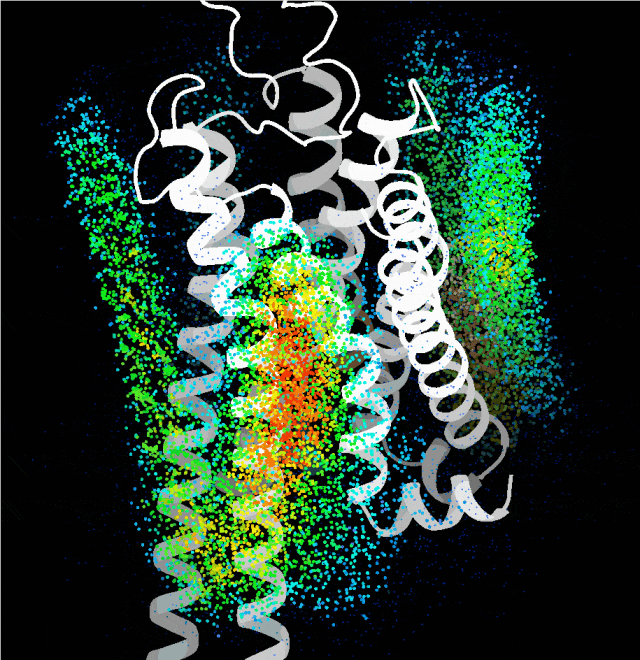
Point cloud plot revealing the most plausible regions for the transmembrane single helix to bind, generated by the probing algorithm.
The team’s idea was to design transmembrane protein partners that bind GPCRs at selected interfaces and stabilize specific conformations, essentially providing a “behavioral constraint” without replacing the receptor’s ability to sense its environment. In their approach, GEMs can also be fused with GPCRs, expanding the classic seven-pass architecture to nine or even thirteen transmembrane segments. The added helices act as a structural framework that can bias how signals are transmitted.
Creating such membrane proteins from scratch is notoriously difficult. De novo design requires generating sequences and folds without relying on natural templates, and for GPCRs the challenge is compounded by the moving parts: the designers must understand which transmembrane contacts participate in the receptor’s mechanical motion and which are most likely to redirect signaling.
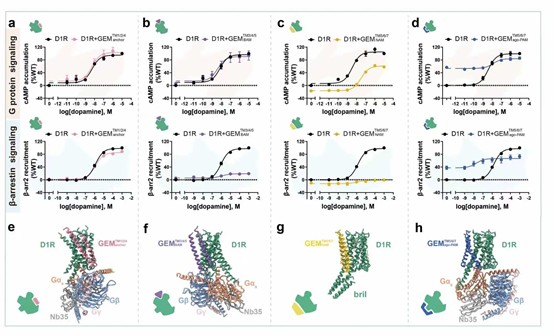
High-resolution structures of de novo designed GEMs in complex with D1R.
To tackle this challenge, the team first built computational “probes” to scan GPCR transmembrane surfaces, especially regions that small molecules struggle to reach, mapping candidate regulatory interfaces. They then combined generative methods, including diffusion models, with a “structural prompting” stragegy that guides the AI toward designs expected to bind specific epitopes and lock in defined conformations. The resulting candidates were tested in cell assays and validated structurally using cryo-electron microscopy.
By iterating between design and experiment, the researchers developed three prompting strategies — site-directed insertion, pre-occupancy blocking and conformation induction — that yielded GEM components capable of distinct behaviors: activating receptors, inhibiting them, or biasing signaling toward particular pathways.
The authors report close consistency between predicted and measured structures, with an accuracy approaching 1.0 Å — an atom-scale match that suggests the design pipeline can reliably build membrane-embedded interfaces rather than merely approximate them. “We’re not just turning receptors on or off,” says ZHANG Min. “The system offers a degree of programmability.” In principle, modular GEM parts could be assembled in circuit-like combinations to implement biological “logic gates” that re-route cellular signals.
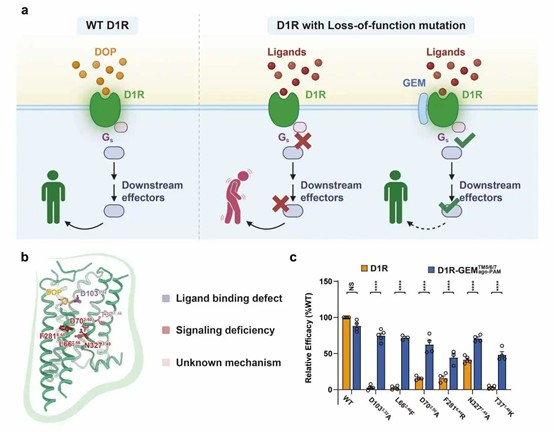
GEM ago-PAM can rescue GPCR Loss-of-function mutants.
ZHANG Yan uses a cultural metaphor to describe the concept. If GPCRs are like Sun Wukong — the Monkey King, famed for his ability to take on hundreds of forms — then GEMs resemble the tightening headband that can rein in his movements. The engineered scaffold is not part of the receptor itself. Instead, it sits at a critical position on the transmembrane surface, guiding, limiting or redirecting the receptor’s conformational repertoire. That kind of control could be particularly valuable for mutation-driven GPCR dysfunctions where receptors can slip off track, or where excessive basal activity calls for soft restraint rather than complete shutdown.
The work also suggests a shift in how researchers might modulate GPCRs. Rather than competing for the orthosteric pocket, designers can create a transmembrane partner that tunes receptor mechanics from the outside. “We hope this approach can offer new therapeutic ideas for GPCR dysfunction diseases that have long lacked effective interventions, for example, certain Parkinson’s disease,” says ZHANG Yan.
In that sense, GEMs point to a new route that differs from traditional small-molecule drugs. BY imposing a programmable “behavioral constraint”, they allow receptors to retain their original sensing ability while responding in new ways. Meanwhile, the study maps out an AI-driven platform for functional design of membrane proteins, offering a fresh perspective for understanding and leveraging life’s signaling systems.
Translator: FANG Fumin
Editor: HAN XIao





