Research Team Led by Prof. Zhang Mingfang Reveals Differential Homoeolog Gene Expression and Selection Behavior
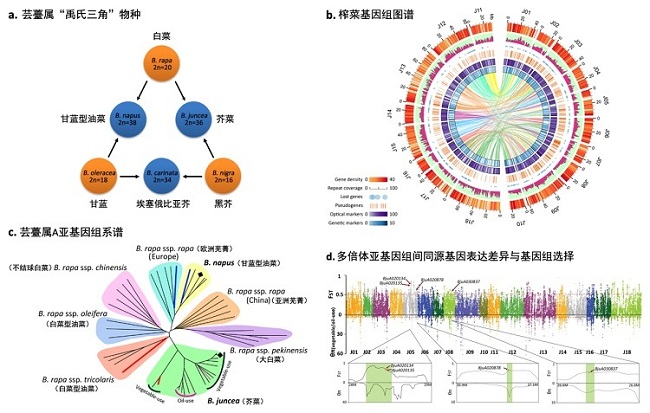
Polyploidization is a prevalent phenomenon during the course of evolution in the plant kingdom. Particularly commonplace in flowering plants, polyploidy is perceived as a defining characteristic of all angiosperm genomes during their evolution and plays an essential role in speciation and genomic plasticity. The Brassica genus contains a diverse range of oilseed and vegetable crops important for human nutrition. Crops of particular agricultural importance include three diploid species, Brassica rapa (AA), Brassica nigra (BB) and Brassica oleracea (CC), and three allopolyploid species, B. napus (AACC), B. juncea (AABB) and Brassica carinata (BBCC). The evolutionary relationships among these Brassica species are described by what is called the ‘triangle of U’ model, which proposes how the genomes of the three ancestral Brassica species, B. rapa, B. nigra and Brassica oleracae, combined to give rise to the allopolyploid species of this genus. B. juncea formed by hybridization between the diploid ancestors of B. rapa and B. nigra, followed by spontaneous chromosome doubling. Subsequent diversifying selection then gave rise to the vegetable- and oil-use subvarieties of B. juncea. These subvarieties include vegetable and oilseed mustard in China, oilseed crops in India, canola crops in Canada and Australia, and condiment crops in Europe and other regions. Cultivation of B. juncea began in China about 6,000 to 7,000 years ago, and flourished in India from 2,300 BC onward.
The research team led by Prof. Zhang Mingfang assembled an allopolyploid Brassica juncea genome by shotgun and single-molecule reads integrated to genomic and genetic maps. We discovered that the A subgenomes of B. juncea and Brassica napus each had independent origins. Results suggested that A subgenomes of B. juncea were of monophyletic origin and evolved into vegetable-use and oil-use subvarieties. Homoeolog expression dominance occurs between subgenomes of allopolyploid B. juncea, in which differentially expressed genes display more selection potential than neutral genes. Homoeolog expression dominance in B. juncea has facilitated selection of glucosinolate and lipid metabolism genes in subvarieties used as vegetables and for oil production. These homoeolog expression dominance relationships among Brassicaceae genomes have contributed to selection response, predicting the directional effects of selection in a polyploid crop genome.
Relevant findings were published online in an article titled “The genome sequence of allopolyploid Brassica juncea and analysis of differential homoeolog gene expression influencing selection” in Nature Genetics. This research has immense significance in the field of allopolyploid evolution and agricultural selection behavior. Experts point out that these homoeolog genes may be contributing to novel features of polyploids and may also be preferential targets for crop improvement.
The multiuse allopolyploid B. juncea genome offers a distinctive model to study the underlying genomic basis for selection in breeding improvement. These findings place this work into the broader context of plant breeding, highlighting a potential link between homoeolog expression dominance and trait improvement that might be extendable to other polyploid crops.