From the evolution of MCR-4 to the functional unification of the MCR family
The research team led by FENG Youjun at the School of Medicine has made major breakthroughs in research into the mechanism behind mobile colistin resistance (MCR) recently. Their findings were published in two articles in Advanced Science on April 3, 2019.
Antimicrobial resistance (AMR) has become one of the biggest threats to global health, food security, and social development, particularly in developing countries, including China and India. The emergence and expeditious spread of a growing body of MCR family enzymes is jeopardizing the renewed interest in colistin as one of the limited options in the clinical sector. FENG Youjun et al. actively engaged in this field and developed a novel perspective on the evolutionary diversification and the functional unification of the MCR family.
In the article entitled “Definition of a Family of Non-mobile Colistin Resistance (NMCR-1) Determinants Suggests Aquatic Reservoirs for MCR-4”, FENG Youjun et al. report a new family of nonmobile colistin resistance (from nmcr‐1 to nmcr‐1.8) from the aquatic bacterium Shewanella. NMCR-1 (541aa) displays 62.78% identity to MCR-4. Genetic and structural analyses reveal that NMCR‐1 shares a similar catalytic mechanism and functional motifs, both of which are required for MCR action and its resultant phenotypic resistance to polymyxin. Phylogeny and domain‐swapping demonstrate that NMCR‐1 is a progenitor of MCR‐4 rather than MCR‐1/2. Additionally, the experiment of bacterial growth and viability reveals that NMCR‐1 promotes fitness cost as MCR‐1/4 does in the recipient Escherichia coli. This finding suggests that the aquatic bacterium Shewanella (and even its associated aquaculture) is a reservoir for MCR‐4 mobile colistin resistance.
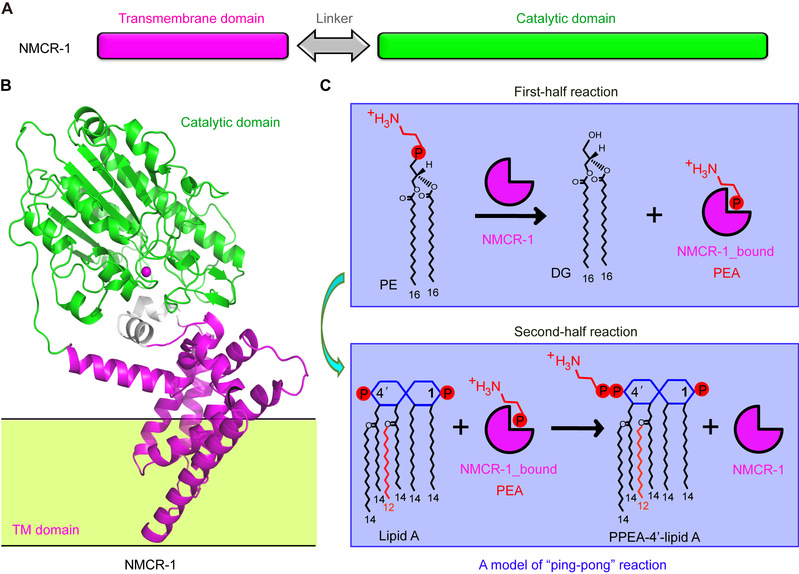
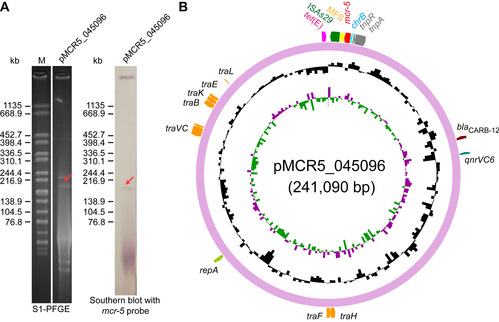
In the article entitled “A Genomic, Evolutionary, and Mechanistic Study of MCR-5 Action Suggests Functional Unification across the MCR Family of Colistin Resistance”, FENG Youjun et al. report the comparative genomics of a large plasmid harboring mcr - 5 from Aeromonashydrophila and the structural/functional perspectives of MCR‐5 action.Whole genome sequencing and comparative genomics have identified the loss of certain parts of the Tn3‐type transposon typically associated with mcr‐5, providing a clue toward its mobilization. Phylogeny of MCR‐5 implies that it is distinct from the MCR‐1/2 sub‐lineage, but it might share a common ancestor of MCR‐3/4. Domain‐swapping analysis of MCR‐5 elucidates that its two structural motifs (transmembrane domain and catalytic domain) are incompatible with its counterparts in MCR‐1/2. Like the rest of the MCR family, MCR‐5 exhibits a series of conservative features, including zinc‐dependent active sites, phosphatidylethanolamine‐binding cavity, and the mechanism of enzymatic action. The study reveals that MCR family resistance enzymes are functionally unified, though taking varied evolutionary pathways.
Given detailed perspectives from genomic, evolutionary, structural, and mechanistic studies, FENG Youjun et al. take a closer step towards developing novel therapeutic agents and adjuvants that can address the whole MCR family.





